Output of Trinity Assembly
When Trinity completes, it will create a 'trinity_out_dir.Trinity.fasta' output file (or prefix based on the output directory you specify).
Trinity groups transcripts into clusters based on shared sequence content. Such a transcript cluster is very loosely referred to as a 'gene'. This information is encoded in the Trinity fasta accession. An example Fasta entry for one of the transcripts is formatted like so:
>TRINITY_DN1000_c115_g5_i1 len=247 path=[31015:0-148 23018:149-246]
AATCTTTTTTGGTATTGGCAGTACTGTGCTCTGGGTAGTGATTAGGGCAAAAGAAGACAC
ACAATAAAGAACCAGGTGTTAGACGTCAGCAAGTCAAGGCCTTGGTTCTCAGCAGACAGA
AGACAGCCCTTCTCAATCCTCATCCCTTCCCTGAACAGACATGTCTTCTGCAAGCTTCTC
CAAGTCAGTTGTTCACAGGAACATCATCAGAATAAATTTGAAATTATGATTAGTATCTGA
TAAAGCA
The accession encodes the Trinity 'gene' and 'isoform' information. In the example above, the accession 'TRINITY_DN1000_c115_g5_i1' indicates Trinity read cluster 'TRINITY_DN1000_c115', gene 'g5', and isoform 'i1'. Because a given run of trinity involves many many clusters of reads, each of which are assembled separately, and because the 'gene' numberings are unique within a given processed read cluster, the 'gene' identifier should be considered an aggregate of the read cluster and corresponding gene identifier, which in this case would be 'TRINITY_DN1000_c115_g5'.
So, in summary, the above example corresponds to 'gene id: TRINITY_DN1000_c115_g5' encoding 'isoform id: TRINITY_DN1000_c115_g5_i1'.
The Path information stored in the header ("path=[31015:0-148 23018:149-246]") indicates the path traversed in the Trinity compacted de Bruijn graph to construct that transcript. In this case, node '31015' corresponds to sequence range 0-148 of the transcript, and node 23018 corresponds to sequence range 149-246 of the transcript sequence. The node numbers are unique only in the context of a given Trinity gene identifier, and so graph nodes can be compared among isoforms to identify unique and shared sequences of each isoform of a given gene.
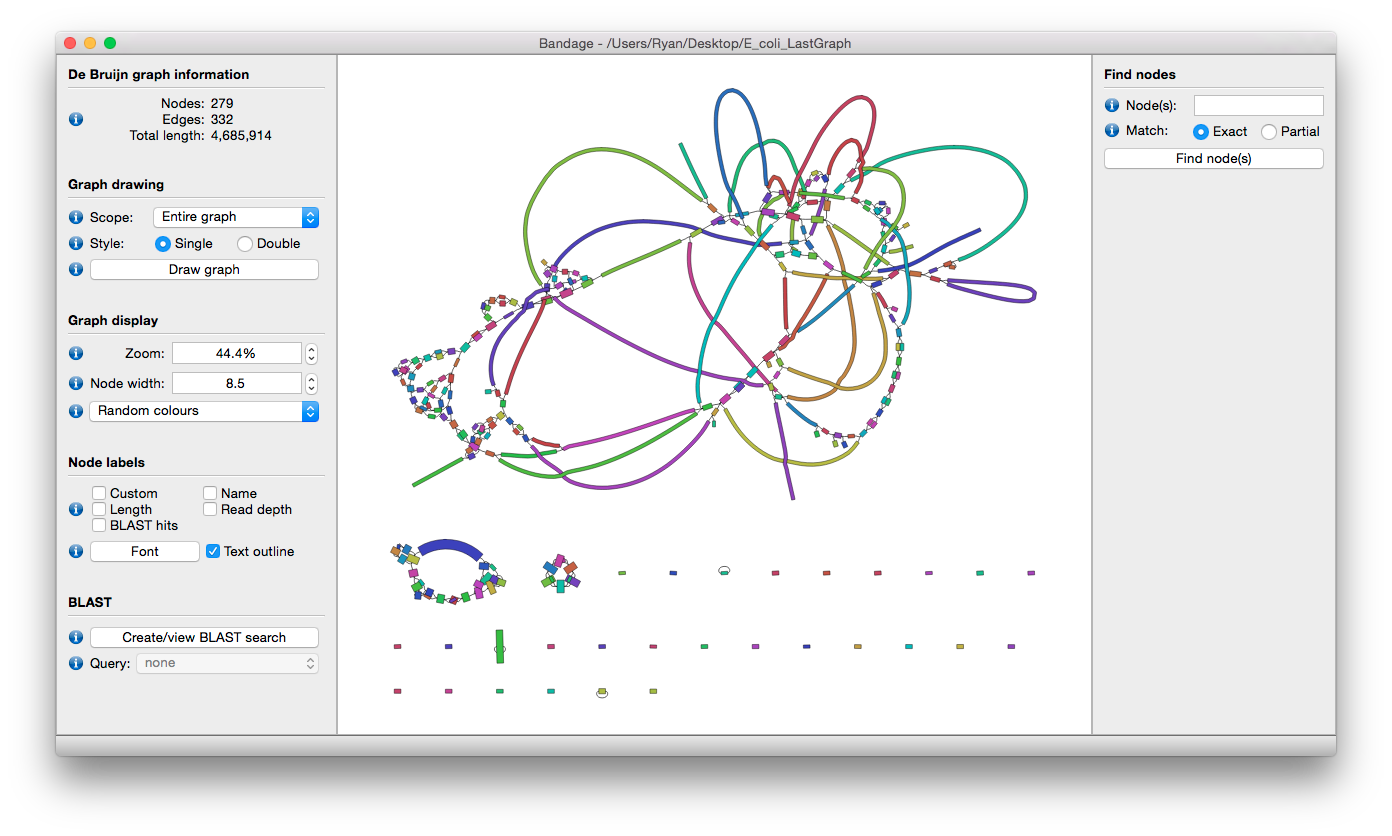
The Trinity assembly can be visualized using Bandage. See video https://www.youtube.com/watch?v=VuRN28XyFcI.
- Trinity Wiki Home
- Installing Trinity
- Running Trinity
- Trinity process and resource monitoring
- Output of Trinity Assembly
- Assembly Quality Assessment
- Downstream Analyses
- Miscellaneous additional functionality that may be of interest
- Contributing code
- Trinity Tidbits
- Frequently Asked Questions (FAQ)
- There are too many transcripts! What do I do?
- How to minimize RAM usage
- How do I use reads I downloaded from SRA
- How do I identify the specific reads that were incorporated into the transcript assemblies?
- How can I perform cross-species analysis?
- How do I combine PE and SE reads?
- How can I run this in parallel on a computing grid?
- Computing and Time requirements
- Errors during Trinity run
- Killing Trinity
- Contact us