Please first read the paper Zhu, Wentao, Yufang Huang, Liang Zeng, Xuming Chen, Yong Liu, Zhen Qian, Nan Du, Wei Fan, and Xiaohui Xie. "AnatomyNet: Deep Learning for Fast and Fully Automated Whole‐volume Segmentation of Head and Neck Anatomy." Medical physics (2018).
Use PDDCA18 (MICCAI) training + HNPETCT + HNCetuximab as training set. Use PDDCA18 test as test.
The data spliting is in https://www.google.com/search?q=PDDCA+dataset&oq=PDDCA+dataset&aqs=chrome..69i57j0l5.3910j0j7&sourceid=chrome&ie=UTF-8
./data/*.csv files are used to remove non-head and neck regions. Processed PDDCA, PETCT and CET datasets can be downloaded from https://drive.google.com/open?id=1yTarwj5_rp7eX2gCm5k8R0RsZVGzdYlc https://drive.google.com/file/d/17SUj2Si3eD8BHQ7nGpPSO_6wSLlk0TEg/view?usp=sharing https://drive.google.com/file/d/1XbsHVVtap72qp11Pv2cUzbV3dosOjBos/view?usp=sharing
The numpy and cropping information in .csv are from original .nrrd data. It is recommended you extract spacing from .nrrd and resample the numpy (z, y, x) to data of spacing 0.76 (x), 0.76 (y), and 1.25 (z) which are the smallest spacing of the dataset if you want to resample to a fixed spacing.
For the files in ./src/, please read the paper for details.
Depedencies: pytorch 0.3/0.4, simpleitk 1.1.0, tqdm 4.11.2, cv2 3.4.0. I recommend to use Anaconda to configure the environment.
preprocee_crop.ipynb can be used for cropping.
baseline.py is the U-Net with only one downsample layer.
baseline2Pool.py is the U-Net with two downsampling layer.
baseline3Pool.py is the U-Net with two downsampling layer.
baseline4Pool.py is the U-Net with two downsampling layer.
baselineDiceCrossEntropy.py is the hybrid loss with dice loss and cross entropy loss.
baselineDiceFocalLoss.py is the hybrid loss with dice loss and focal loss.
baselineExpLogLoss.py is the exponential log loss. It does not work well both theoretically and experimentally. You can read the paper for detail.
baselineRes18Conc.py uses Res18 as the encoder and concatenation in the skip connection.
baselineRes18Sum.py uses Res18 as the encoder and summation in the skip connection.
baselineSERes18Conc.py uses Squeeze and Excitation Res18 as the encoder and concatenation in the skip connection.
baselineSERes18sum.py uses Squeeze and Excitation Res18 as the encoder and summation in the skip connection.
AnatomyNet.py is used for the proposed method with the initialization of baselineSERes18Conc.py
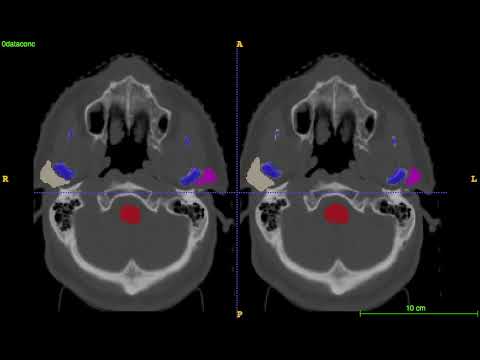
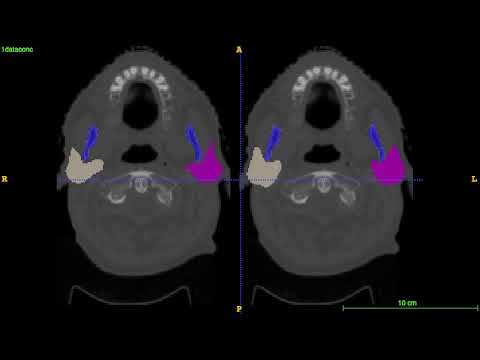
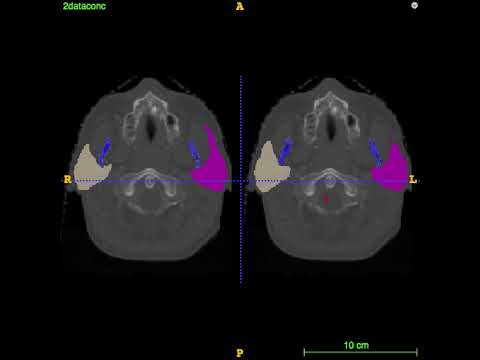
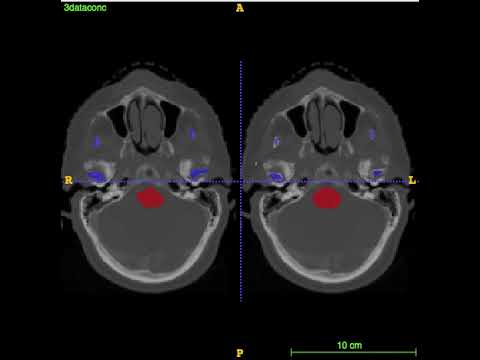
Segmentation for the first 4 test CT images on MICCAI 2015 challenge: left is the ground truth; right is the prediction.