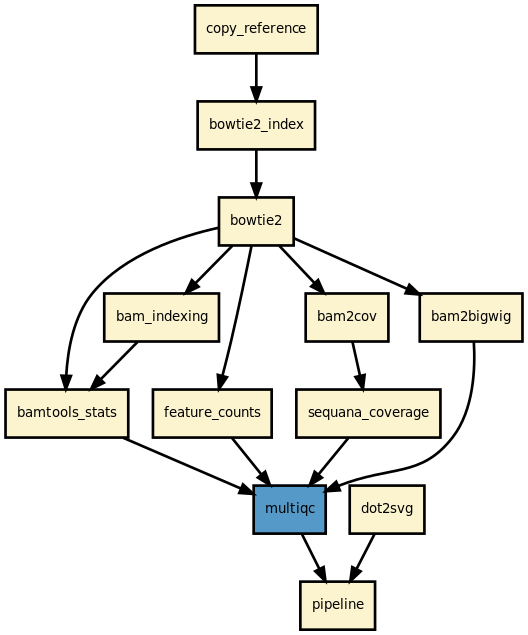
This is the mapper pipeline from the Sequana projet
- Overview
This is a simple pipeline to map several FastQ files onto a reference using different mappers/aligners
- Input
A set of FastQ files (illumina, pacbio, etc).
- Output
A set of BAM files (and/or bigwig) and HTML report
- Status
Production
- Documentation
This README file, and https://sequana.readthedocs.io
- Citation
Cokelaer et al, (2017), 'Sequana': a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI https://doi:10.21105/joss.00352
If you already have all requirements, you can install the packages using pip:
pip install sequana_mapper --upgradeYou will need third-party software such as fastqc. Please see below for details.
This command will scan all files ending in .fastq.gz found in the local directory, create a directory called mapper/ where a snakemake pipeline can be executed.:
sequana_mapper --input-directory DATAPATH --mapper bwa --create-bigwig
sequana_mapper --input-directory DATAPATH --mapper bwa --do-coverageThis creates a directory with the pipeline and configuration file. You will then need to execute the pipeline:
cd mapper
sh mapper.sh # for a local runThis launch a snakemake pipeline. If you are familiar with snakemake, you can retrieve the pipeline itself and its configuration files and then execute the pipeline yourself with specific parameters:
snakemake -s mapper.rules -c config.yaml --cores 4 \
--wrapper-prefix https://raw.githubusercontent.com/sequana/sequana-wrappers/Or use sequanix interface.
This pipelines requires the following executable(s):
- bamtools
- bwa
- multiqc
- sequana_coverage
- minimap2
- bowtie2
- deeptools
This pipeline runs mapper in parallel on the input fastq files (paired or not). A brief sequana summary report is also produced. When using --pacbio option, -x map-pb options is automatically added to the config.yaml file and the readtag is set to None.
The BAM files are filtered to remove unmapped reads to keep BAM files to minimal size. However, the multiqc and statistics to be found in {sample}/bamtools_stats/ includes mapped and unmapped reads information. Each BAM file is stored in a directory named after the sample.
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
| Version | Description |
|---|---|
| 1.2.1 | * fix bwa_split bwa aggreate stage (bug fix) |
1.2.0 |
|
1.1.0 |
|
| 1.0.0 | * Use latest sequana-wrappers and graphviz apptainer |
| 0.12.0 | * Use latest pipetools and add singularity containers |
| 0.11.1 | * Fix typo when setting coverage to True and allow untagged filenames |
| 0.11.0 | * implement feature counts for capture-seq projects |
| 0.10.1 | * remove getlogdir and getname |
| 0.10.0 | * use new wrappers framework |
0.9.0 |
|
| 0.8.13 | * add the thread option in minimap2 case |
| 0.8.12 | * factorise multiqc rule |
0.8.11 |
|
0.8.10 |
|
| 0.8.9 | * fix requirements |
0.8.8 |
|
| 0.8.7 | * fix config file creation (for bigwig) |
| 0.8.6 | * added bowtie2 mapper + bigwig as output, make coverage optional |
| 0.8.5 | * create a sym link to the HTML report. Better post cleaning. |
| 0.8.4 | * Fixing multiqc (synchronized with sequana updates) |
| 0.8.3 | * add sequana_coverage rule. |
| 0.8.2 | * add minimap2 mapper |
| 0.8.1 | * fix bamtools stats rule to have different output name for multiqc |
| 0.8.0 | First release. |
To contribute to this project, please take a look at the Contributing Guidelines first. Please note that this project is released with a Code of Conduct. By contributing to this project, you agree to abide by its terms.