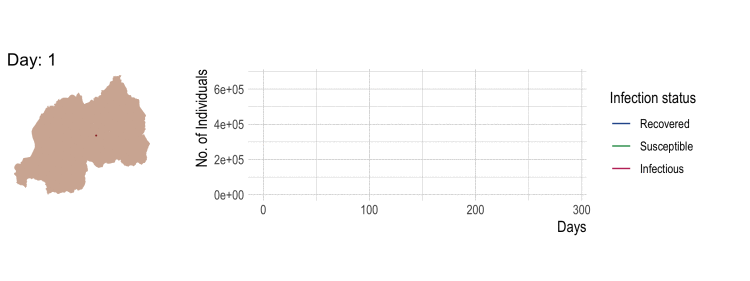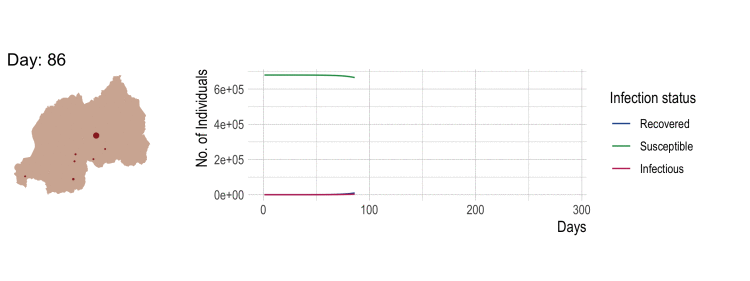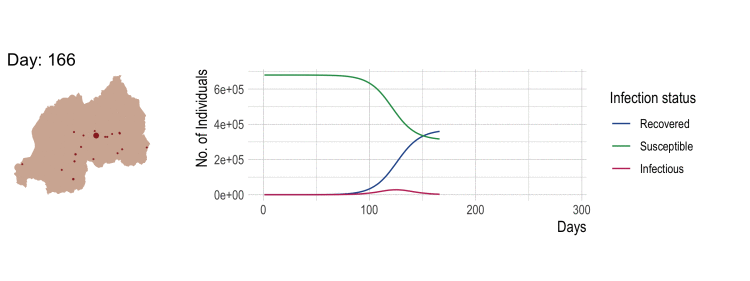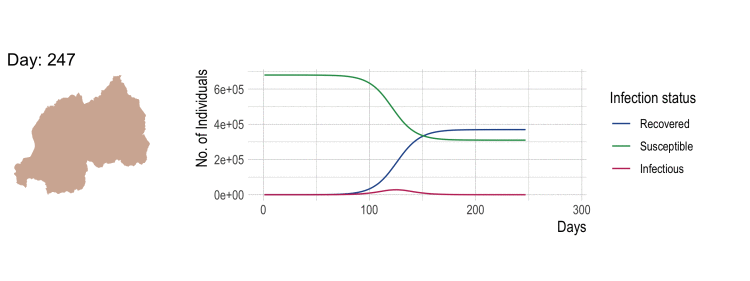
SEEDNet has been designed to simulate outbreaks over a network created by combining satellite imagery, population data, and road network information. It is versatile in that it can be used for any metapopulation model on networks with distance and population data.
Run the following:
remotes::install_github("nistara/SEEDNet")library("SEEDNet")Discrete time, stochastic, metatpopulation model.
In case of influenza, the compartmental model is SEIR:
S - Susceptible E - Latent I - Infectious R - Recovered
-
disnet_commuting- Comuting function. Takes in network object (ingraphmlformat), and calculates the commuting rates for all outgoing edges in network -
disnet_sim_setup- Simulation setup function. Takes in the graph file with commuting rates added to it, and preps it up for runnning simulations. -
disnet_simulate- Simulation function. This function runs the disease model simulations over the network.
library("disnet")
# Read in sample graph/network
f = system.file("sampleData", "g.rds", package = "disnet")
g = readRDS(f)
# calculate commuting rates over it
g_comm = disnet_commuting(g)
# select random node to seed infection in
set.seed(890)
nodes = igraph::vcount(g_comm)
seed_nd = igraph::vertex_attr(g_comm, "name", sample(1:nodes, 1))
# set up the network for simulations
for_sim = disnet_sim_setup(g_comm, seed_nd = seed_nd, output_dir = NA)
# run the simulations over the network
simres = disnet_simulate(sim_input = for_sim, sim_output_dir = NA)g.RDS: The raw graphml/network object
To access the sample dataset:
f = system.file("sampleData", "g.rds", package = "disnet")
g = readRDS(f)If you would like to contribute to this software, please submit a pull request to the GitHub repository. Feel free to report issues, provide feedback, suggest improvements, and ask questions by opening a new issue.