-
Notifications
You must be signed in to change notification settings - Fork 0
Home
Welcome to the sequencingData_track_R wiki!
The R code was developed to draw the visualization of signals, which were produced from the next-generation sequencing data, such as Chip-seq, RNA-seq, BS-seq, and so on. In particular, the preprocessed data is a normalized signals via the function 'genomeCov' of bedtools. After that, the R code may be called to draw signal tracking for a batch of samples. For more details, please check README.
Reference: Jianlin He, Xiguang Xu, Aboozar Monavarfeshani, Sharmi Banerjee, Michael A. Fox, Hehuang Xie. Retinal-input-induced epigenetic dynamics in the developing mouse dorsal lateral geniculate nucleus.Epigenetics & Chromatin (2019) 12: 13.
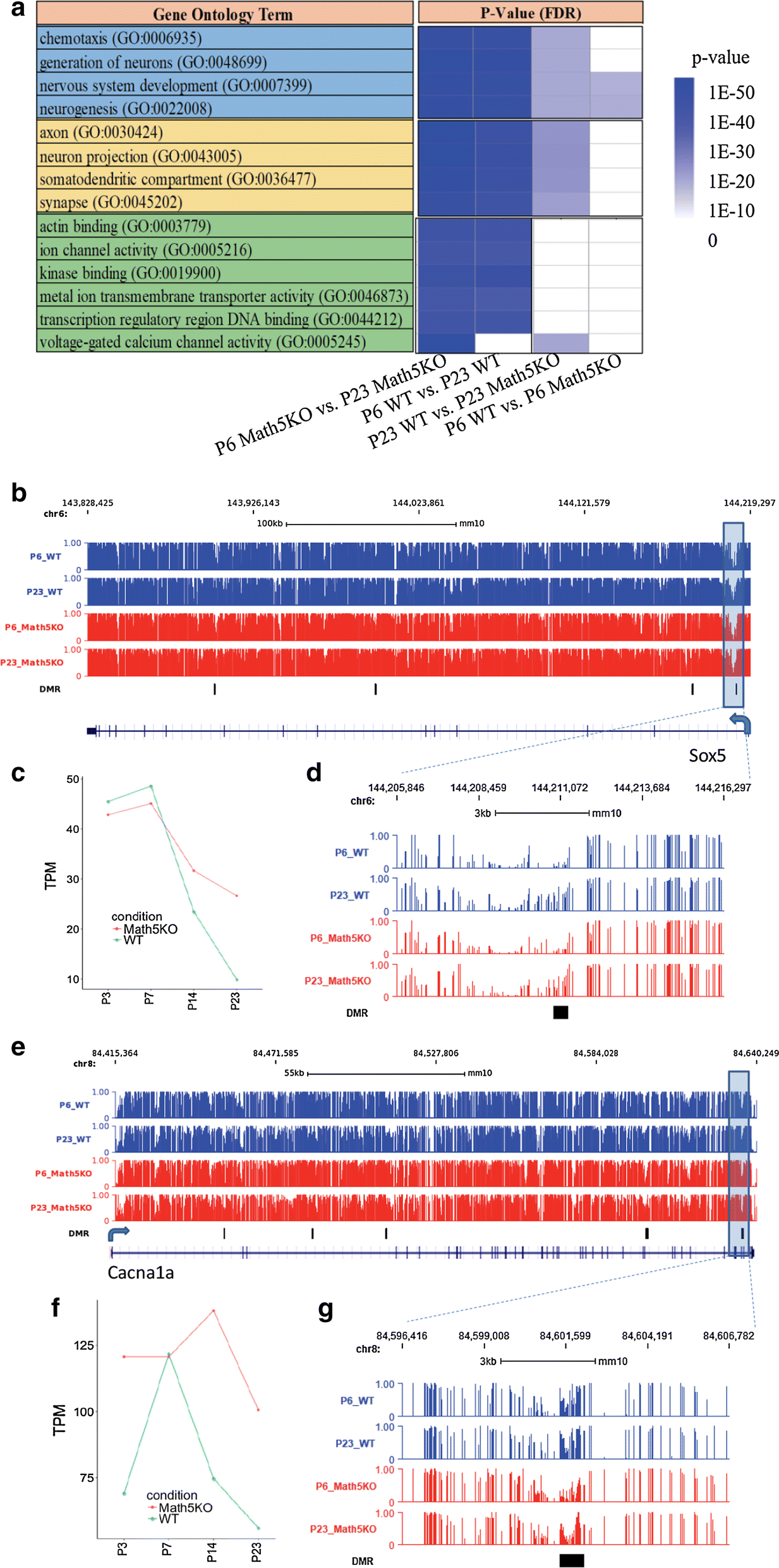
Fig. 4 GO enrichment (a) of genes associated with dLGN epigenetic changes and gene expression and DNA methylation profiles for Sox5 (b–d) and CACNA1A (e–g). The genes associated with dLGN methylation changes are with DMRs in gene body or within 10 kb upstream from their transcriptional starting sites. a The color bar from white to dark blue denotes the significance of GO term from low to high. The genomic loci for b and e are chr6:143,828,425-144,219,297 and chr8:84,415,364-84,640,249, respectively