19-11-2021 Update: Bug fixes, ability to load fasta file with more than one sequence and select tab is provided.
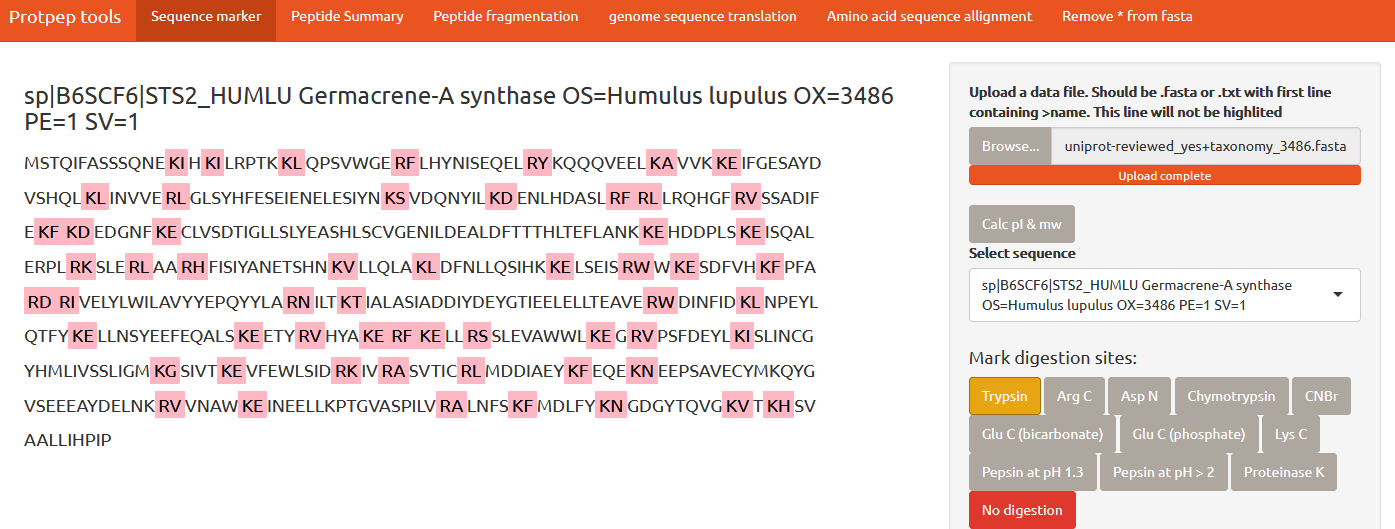 Simple shiny app for protease cleavage sites in proteins visualization. In addition to this, some basic tools for protein/peptide properties calculation and sequence data (pre)processing are provided.
Simple shiny app for protease cleavage sites in proteins visualization. In addition to this, some basic tools for protein/peptide properties calculation and sequence data (pre)processing are provided.
You can download attached script and run your app locally or use web based version: https://mukhomorr.shinyapps.io/pep_marker/
Don't forget to change your working directory before run app on your machine.
**This app was made during first attemts of shiny apps writing and do not challenges to be something necessary for serious research,. But as there are people, who don't familliar with various programming languages and there are no freely avalible resource gui based programms for solving that types of task - why not? Despite the fact, that this is only training exersice for myself - updates will be provided.
-
Notifications
You must be signed in to change notification settings - Fork 0
Simple shiny app for protease cleavage sites in proteins visualization. Could be helpfull on hplc-ms/ms based experiment design step. In addition to this, some basic tools for protein/peptide properties calculation and sequence data (pre)processing are provided.
License
mukhomorr/PepShinyToolsR
This commit does not belong to any branch on this repository, and may belong to a fork outside of the repository.
Folders and files
| Name | Name | Last commit message | Last commit date | |
|---|---|---|---|---|
Repository files navigation
About
Simple shiny app for protease cleavage sites in proteins visualization. Could be helpfull on hplc-ms/ms based experiment design step. In addition to this, some basic tools for protein/peptide properties calculation and sequence data (pre)processing are provided.