-
Notifications
You must be signed in to change notification settings - Fork 1.5k
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
feat(bayesian_optimization.py): Allow registering of temporary observations #365
base: master
Are you sure you want to change the base?
Conversation
Codecov ReportBase: 99.25% // Head: 97.16% // Decreases project coverage by
Additional details and impacted files@@ Coverage Diff @@
## master #365 +/- ##
==========================================
- Coverage 99.25% 97.16% -2.09%
==========================================
Files 8 8
Lines 535 564 +29
Branches 81 89 +8
==========================================
+ Hits 531 548 +17
- Misses 2 13 +11
- Partials 2 3 +1
Help us with your feedback. Take ten seconds to tell us how you rate us. Have a feature suggestion? Share it here. ☔ View full report at Codecov. |
|
Hey @Rizhiy, thanks for contributing! I (superficially) follwed the discussion in the original issue and I agree with the notions that a) this package should be compatible with parallel optimization and b) in it's current state, that can't really be considered the state (as you have shown). Nonetheless, I'm always somewhat sceptical of messing with internal data and even moreso here, since it involves providing "wrong" values. Is there some way that this could go wrong? Off the top of my head:
If I understand correctly, the idea behind this technique of registering dummies is to reduce the covariance of the GPR at a point that is being sampled so that other workers are suggested other points. Is there a way to perform this covariance reduction without registering dummies? Alternatively, is there a way to force different points to be returned without using dummies, e.g.
In particular, I think that modifying |
|
Hi @till-m, regarding your concerns:
Not sure what you mean, dummy points are stored in Interactions for other observations should be the same, can you perhaps try it out and I can correct the wrong behaviour.
They persist until you overwrite them.
There very well might be, but I haven't dived deep into how the optimiser works on the inside yet.
I will look into that. |
|
Hi @Rizhiy, I misread the code and after thinking about it again, I think you've covered the scenario that I was picturing. My apologies. |
a0e9e50
to
94cd1a2
Compare
|
@till-m I have looked into other ways you suggested achieving similar effect and currently it seems that they won't be as effective or too difficult to implement. There very well might be a better approach for parallel evaluation, but I haven't been able to find any yet. |
|
Looking more closely at the code, perhaps it would make more sense to put dummy logic into |
|
Hi @Rizhiy,
I agree. I think logging should also be considered -- dummies probably shouldn't show up in logs, especially when the
Agreed.
Thanks for looking into this. I also had a look around the literature and I found a few potentially interesting things. Your approach seems to be related to "Constant Liars", see Ginsbourger et al; 2008. Consider also the "Kriging Believer" which may be a more natural way to pick the values of the dummies. I would also like to highlight the approach outlined in Batch Bayesian Optimization via Local Penalization (see also here), that works by penalizing the acquisition function and would be close to what I was thinking of. Finally, Kandasamy et al might also be worth considering, since it seems to work very well while not being too hard to implement, at least at first glance. There's a lot more literature on this problem than I thought there would be, two others that I found potentially interesting would be:
|
|
Hi @Rizhiy, I decided to write a quick implementation of Kandasamy to see it work. The preliminary results are pretty disappointing, will post later. |
|
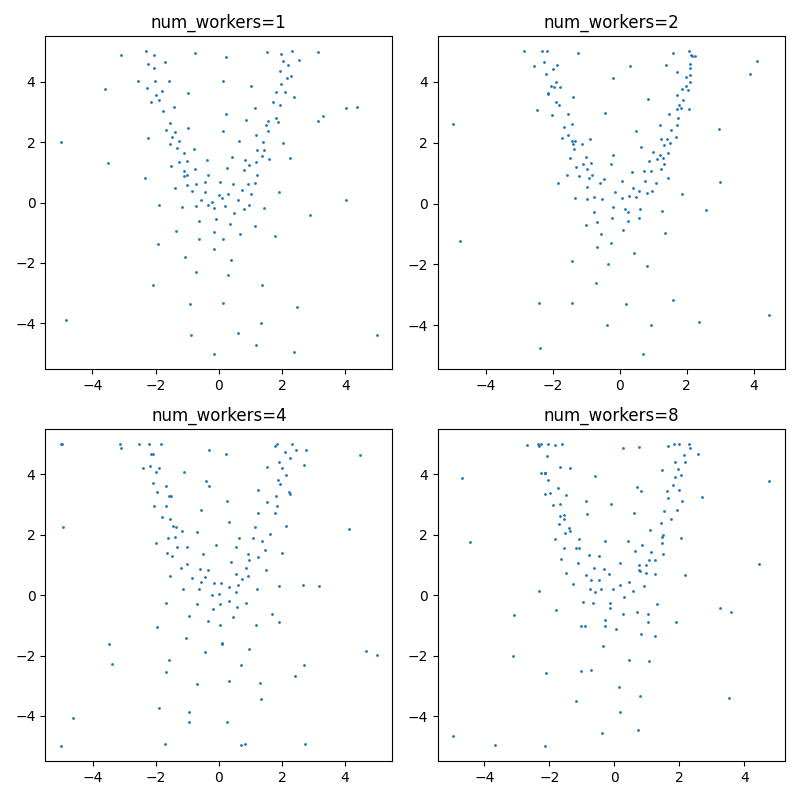
Okay, so first of all it looks like sampling from the multivariate normal is very computationally expensive. I had to adapt Output: I'm somewhat disappointed with these results. It doesn't really seem to hone in on the max very well. Additionally, sampling from the multivariate normal also seems to be quite computationally intensive -- I takes around 4x as long to run the script with Output ( from __future__ import annotations
from typing import Callable
import matplotlib.pyplot as plt
import numpy as np
from bayes_opt.bayesian_optimization import BayesianOptimization
from bayes_opt.util import UtilityFunction
from pandas.core.generic import pickle
from scipy.optimize import rosen
from scipy.stats import multivariate_normal
from tqdm import tqdm
from sklearn.gaussian_process import GaussianProcessRegressor
import warnings
class asyTS():
def __init__(self) -> None:
self.random_state_generator = lambda: np.random.randint(0, 100000)
@property # I know that this is giga-hacky monkey patching, please bear with me.
def utility(self):
random_state = self.random_state_generator()
def func(x, gp: GaussianProcessRegressor, y_max):
with warnings.catch_warnings():
warnings.simplefilter("ignore")
# fixing random state causes it to return the same function
return gp.sample_y(x, random_state=random_state)
return func
def _closest_distance(point, points):
return min(np.linalg.norm(point - p) for p in points if p is not point)
def optimize(
func: Callable[..., float], num_iter: int, bounds: dict[str, tuple[float, float]], num_workers=0, async_thompson=True
):
init_samples = int(np.sqrt(num_iter))
optimizer = BayesianOptimization(f=None, pbounds=bounds, verbose=0)
init_kappa = 10
kappa_decay = (0.1 / init_kappa) ** (1 / (num_iter - init_samples))
if async_thompson:
utility = asyTS()
else:
utility = UtilityFunction(
kind="ucb", kappa=init_kappa, xi=0.0, kappa_decay=kappa_decay, kappa_decay_delay=init_samples
)
init_queue = [optimizer.suggest(utility) for _ in range(init_samples)]
result_queue = []
tbar = tqdm(total=num_iter, leave=False)
while len(optimizer.res) < num_iter:
sample = init_queue.pop(0) if init_queue else optimizer.suggest(utility)
loss = func(list(sample.values())) * -1
result_queue.append((sample, loss))
if len(result_queue) >= num_workers:
try:
optimizer.register(*result_queue.pop(0))
tbar.update()
except KeyError:
pass
return optimizer.res
bounds = {"x": [-5, 5], "y": [-5, 5]}
all_results = {}
for num_workers in tqdm([1, 2, 4, 8], desc="Checking num_workers"):
results = []
results = optimize(rosen, 150, bounds, num_workers, async_thompson=True)
all_results[num_workers] = results
with open("results.pkl", "wb") as f:
pickle.dump(all_results, f)
with open("results.pkl", "rb") as f:
all_results = pickle.load(f)
fig, axs = plt.subplots(2, 2)
fig.set_figheight(8)
fig.set_figwidth(8)
axs = [item for sublist in axs for item in sublist]
for idx, (num_workers, results) in enumerate(all_results.items()):
if num_workers > 8:
continue
samples = [np.array(list(res["params"].values())) for res in results]
axs[idx].scatter(*zip(*samples), s=1)
axs[idx].set_title(f"{num_workers=}")
avg_min_distance = np.mean([_closest_distance(sample, samples) for sample in samples])
best_result = max([res["target"] for res in results])
print(f"{num_workers=}, mean_min_distance={avg_min_distance:.3f}, {best_result=:.3f}")
fig.tight_layout()
fig.savefig("results.png") |
|
Is there something similar to kappa decay in your code? |
|
My apologies for the delay.
In theory this shouldn't be necessary, but I could add it. I will try to get around to doing that. Overall, I'm not sure if the TS approach is worth exploring due to the sampling speed. |
|
Hi @Rizhiy, apologies for letting this sit for a while. I've been thinking and we should definitely have functionality for this, but it would be better to first redesign the acquisition function API. I've implemented some ideas and will push them shortly, including an implementation of the Kriging believer algorithm. Would you be interested in reviewing and/or testing that code? |
|
Hi, I can go over it briefly, but TBH I haven't been looking in this area since last year, since my approach works fine for me. |
Potential fix for #347