Multidimensional diffusion MRI (MD-dMRI) is a family of conceptually related methods relying on advanced gradient modulation schemes and data processing approaches to simultaneously quantify several microstructural and dynamical properties of tissue by separating their effects on the detected MRI signal into multiple acquisition and analysis dimensions.
This GitHub repository mainly contains MATLAB software for analysis of MD-dMRI data, and also some auxiliary routines for setup of acquisition protocols, motion correction, and image visualization.
Three families of methods are currently implemented within the framework of MD-dMRI:
- Diffusion tensor distributions
- Diffusional exchange
- Diffusion and incoherent flow
The methods are described briefly here, in some more detail in two review articles,1,2 and more exhaustively in the original publications cited for each method.
Run setup_paths in the root folder to put files in the Matlab path.
To analyze data, you can use the method_name_4d_data2fit and method_name_4d_fit2param functions, where method_name refers to one of the methods implemented here. Alternatively, you can use the mdm_fit command. See mdm_fit --help for an overview of how to use the command. An example is provided below:
mdm_fit --data input/data.nii.gz ...
--mask tmp/mask.nii.gz ...
--method dtd_codivide ...
--out tmp/ ...
--xps input/data_xps.mat;The --data argument should point to a nifti file. The --mask argument is optional, but supplying e.g a brain mask may speed up the analysis by avoiding analysis of background voxels. For an overview of potential --method arguments, see the methods folder. The --out argument takes a folder/prefix as input. Finally, you need to supply data on how the experiment was performed, by giving either the --xps flag with the argument pointing to a .mat file holding an 'eXperiment Parameter Structure' (xps). Alternatively, for diffusion tensor distribution (dtd) methods, you can supply a b-tensor file as an argument to the --btens flag.
We assume that the data has already been preprocessed before this function is called. Preprocessing should include motion and eddy current correction and potentially smoothing. These steps are also supported by the framework (see below).
Construction of the xps or the b-tensor input can be challenging. Some help is provided below.
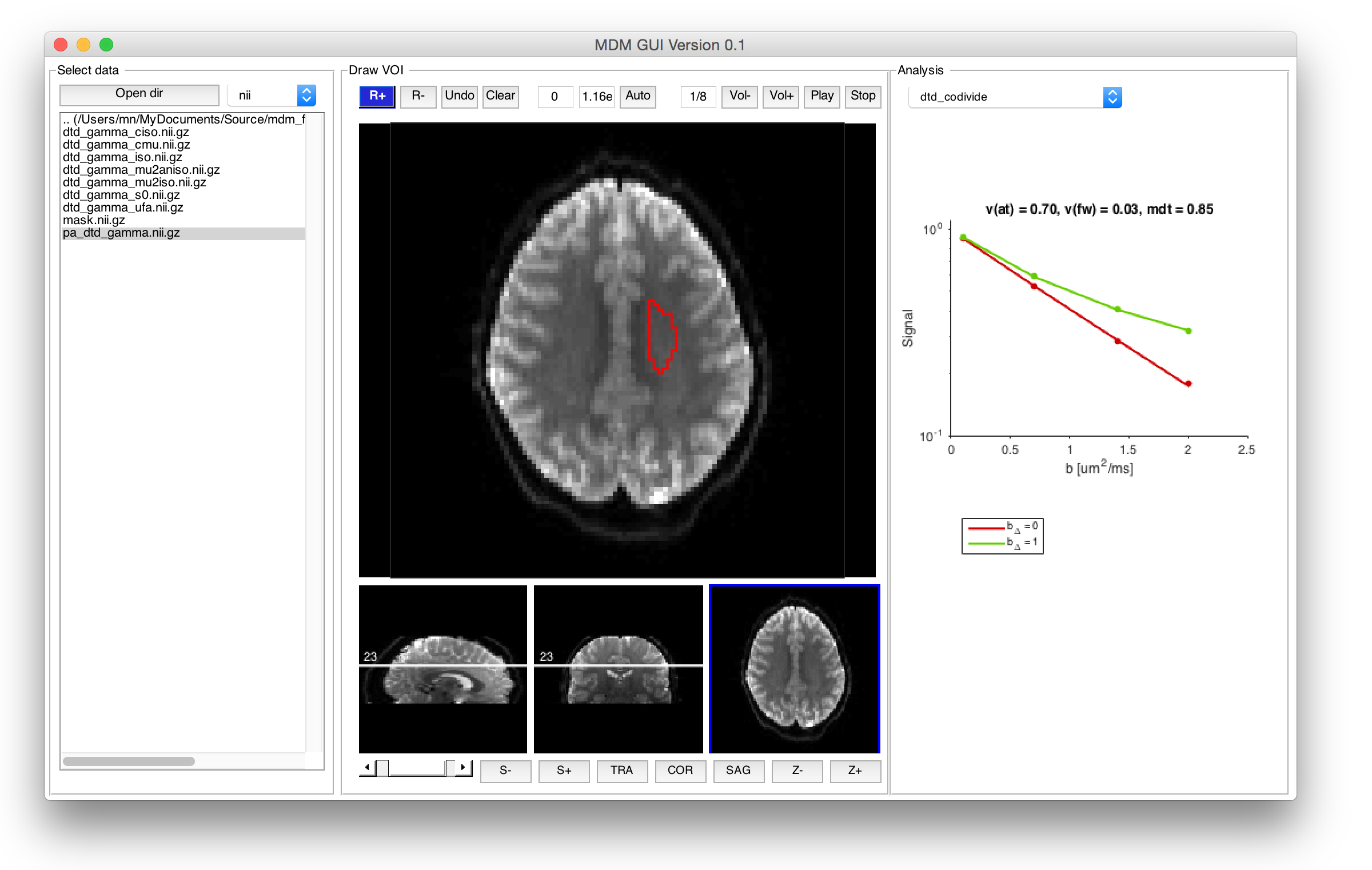
The function mgui starts a graphical user interface that help reviewing the data quality. The left panel shows a folder structure. When opening a data file named e.g. data.nii.gz and the GUI finds an .mat file named as data_xps.mat, it will automatically load it. Then you can draw an ROI an select different methods for fitting the signal data from the dropdown menu in the 'analysis' panel to the right.
Conventional motion correction where all data is registered to a volume acquired with b = 0 s/mm2 is implemented in mdm_mec_b0. This type of correction is not appropriate for high b-value data, where other approached such as extrapolation-based registration should be applied (see this paper). This is supported in the mdm_mec_eb method. For an example, see below
% Options
opt = mdm_opt();
% Connect to the data
s.nii_fn = fullfile(pwd, 'data.nii');
s.xps = mdm_xps_from_bval_bvec('data.bval', 'data.bvec');
% Determine output path
o_path = pwd;
% Write the elastix parameter file
p = elastix_p_affine(200);
p_fn = elastix_p_write(p, 'p.txt');
% Run an extrapolation-based registration
s_registered = mdm_s_mec(s, p_fn, o_path, opt);Now s_registered can be used in the subsequent analysis.
The xps holds all relevant experimental information, i.e., how the experiment was performed. All variables are given in SI-units, and have predetermined names according to this specification. Please see functions mdm_xps_* for help on how to construct an xps. Here we give two examples. First, in the case where a single nii file is analyzed. Secondly, we show an example of how multiple nii files are merged and analyzed.
Creating an xps based on a single nii file
If a single nii file is the basis for the analysis, a single path is used to define its location s.nii_fn = 'nii_filename.nii';. Each nii file is usually associated with files that contain information about the diffusion encoding directions and b-values that were used. These can be used to automatically generate appropriate xps structures. To do this, call
s.xps = mdm_xps_from_gdir('gdir_filename.txt', b_delta);or
s.xps = mdm_xps_from_bval_bvec('bval_filename.bval', 'bvec_filename.bvec', b_delta);depending on what format is used. Note that the files currently do not contain information about the shape of the b-tensor, thus b_delta must be specified by the user. Furthermore, the path to gdir.txt and .bval/.bvec files can be generated based on nii_fn by calling either mdm_fn_nii2gdir or mdm_fn_nii2bvalbvec. The resulting xps can be saved by using mdm_xps_save. If the name of the .mat-file holding the xps is related to the nii filename as specified in mdm_xps_fn_from_nii_fn, the framework will automatically find the xps.
Example: xps from single nii file
This example code requires only that the user defines the filename of the nii to create and save the corresponding xps. Note that this assumes that the gdir.txt or .bval/.bvec files are in the same folder an have standardized names.
% Define path to nii file
s.nii_ fn = 'path_to_nii.nii';
% We assume that the encoding is linear
b_delta = 1;
% Get path to corresponding gdir, and generate the xps
gdir_fn = mdm_fn_nii2gdir(s.nii_fn, b_delta);
s.xps = mdm_xps_from_gdir(gdir_fn, b_delta);
% Or, get path to corresponding bval/bvec, and generate the xps
[bval_fn, bvec_fn] = mdm_fn_nii2bvalbvec(s.nii_fn);
s.xps = mdm_xps_from_bval_bvec(bval_fn, bvec_fn, b_delta);
% Finally, save the xps.
xps_fn = mdm_xps_fn_from_nii_fn(s.nii_fn);
mdm_xps_save(s.xps, xps_fn)Creating an xps based on multiple nii files
In some cases all the necessary data cannot be acquired in a single series, which results in multiple nii files that need to be combined during the analysis. For example, encoding with linear and spherical b-tensors is currently performed in two separate series, but we wish to analyze them simultaneously. In the case when the analysis relies on multiple nii files we first create partial a cell array of partial s structures, and then call mdm_s_merge to merge them all into one. There are also separate functions to merge nii files and xps structures (mdm_nii_merge and mdm_xps_merge).
Example: xps from multiple nii files
This example uses alla the tools form the previous example. It assumes that full filenames are specified to all necessary nii files, and that corersponding gdir.txt or .bval/.bvec files with standardized names are in the same folder (see mdm_fn_nii2gdir and mdm_fn_nii2bvalbvec for standard names). All necessary nii files and corresponding xps structures are first stored in a cell array of structures. Then the cell array of partial s structures is merged. Note that this example includes only the case where gdir.txt files are used to create the xps, but can be modified to use .bval/.bvec according to the example above.
% Create a cell array of s structures.
s{1}.nii_fn = 'nii_filename_1.nii';
s{2}.nii_fn = 'nii_filename_2.nii';
s{3}.nii_fn = 'nii_filename_3.nii';
% Corresponding b-tensor shapes. In this case: spherical, linear, and planar.
b_deltas = [0 1 -.5];
% Define a name for the merged nii (output)
merged_nii_path = '...\output_directory\';
merged_nii_name = 'merged_nii_name.nii';
% Loop over nii files to create partial xps structures, and store them in the cell array.
for i = 1:numel(file_list)
gdir_fn = mdm_fn_nii2gdir(s{i}.nii_fn);
s{i}.xps = mdm_xps_from_gdir(gdir_fn, [], b_deltas(i));
end
% Merge the s structure, and save the merged nii along with its corresponding xps.mat file.
s_merged = mdm_s_merge(s, merged_nii_path, merged_nii_name);
Get acquainted with by xps structure by reading mdm/readme.txt.An extensive description of the code structure is found at http://markus-nilsson.github.io/md-dmri/.
Markus Nilsson, Filip Szczepankiewicz, Björn Lampinen, André Ahlgren, João P. de Almeida Martins, Samo Lasic, Carl-Fredrik Westin, and Daniel Topgaard. An open-source framework for analysis of multidimensional diffusion MRI data implemented in MATLAB. Proc. Intl. Soc. Mag. Reson. Med. (26), Paris, France, 2018. http://archive.ismrm.org/2018/5355.html
-
methods/dtd_smrCode for fitting the model used in Towards unconstrained compartment modeling in white matter using diffusion-relaxation MRI with tensor-valued diffusion encoding, published 06 March 2020 in MRM. See also https://github.com/belampinen/lampinen_mrm_2019. -
methods/ning18Code for fitting the model used in Manuscript Time dependence in diffusion MRI predicts tissue outcome in ischemic stroke patients submitted to MRM in 2020.