New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Speedups for ft_specest_mtmconvol #2215
Comments
|
Hi, @NirOfir sorry to be so slow on this, but here are my initial thougths: in general I am a sucker about code efficiency myself, but on the other hand sometimes detailed profiling and code optimization to save a few % of computation time might not be worth the hassle :). Anyway, I think in general we have to put up with the time spent in ifft'ing the data (which is about half the time that the function needs). Now, w.r.t. your detailed points: I'd be happy to review a PR based on the above :) |
|
What's the status of this @NirOfir ? |
|
Thanks for the feedback! I'm a bit busy right now, but I hope I can get around to working on a PR in the next month or so. Nothing is too urgent about this issue anyway :) |
Is your feature request related to a problem? Please describe.


A few ideas to speed up computation of TFRs and maybe make its feedback more readable. Here are the results of the matlab profiler on data from a single subject with 60 channels sampled at 512 Hz and 795 trials that are 2 seconds long each:
Looking into ft_specest_mtmconvol:
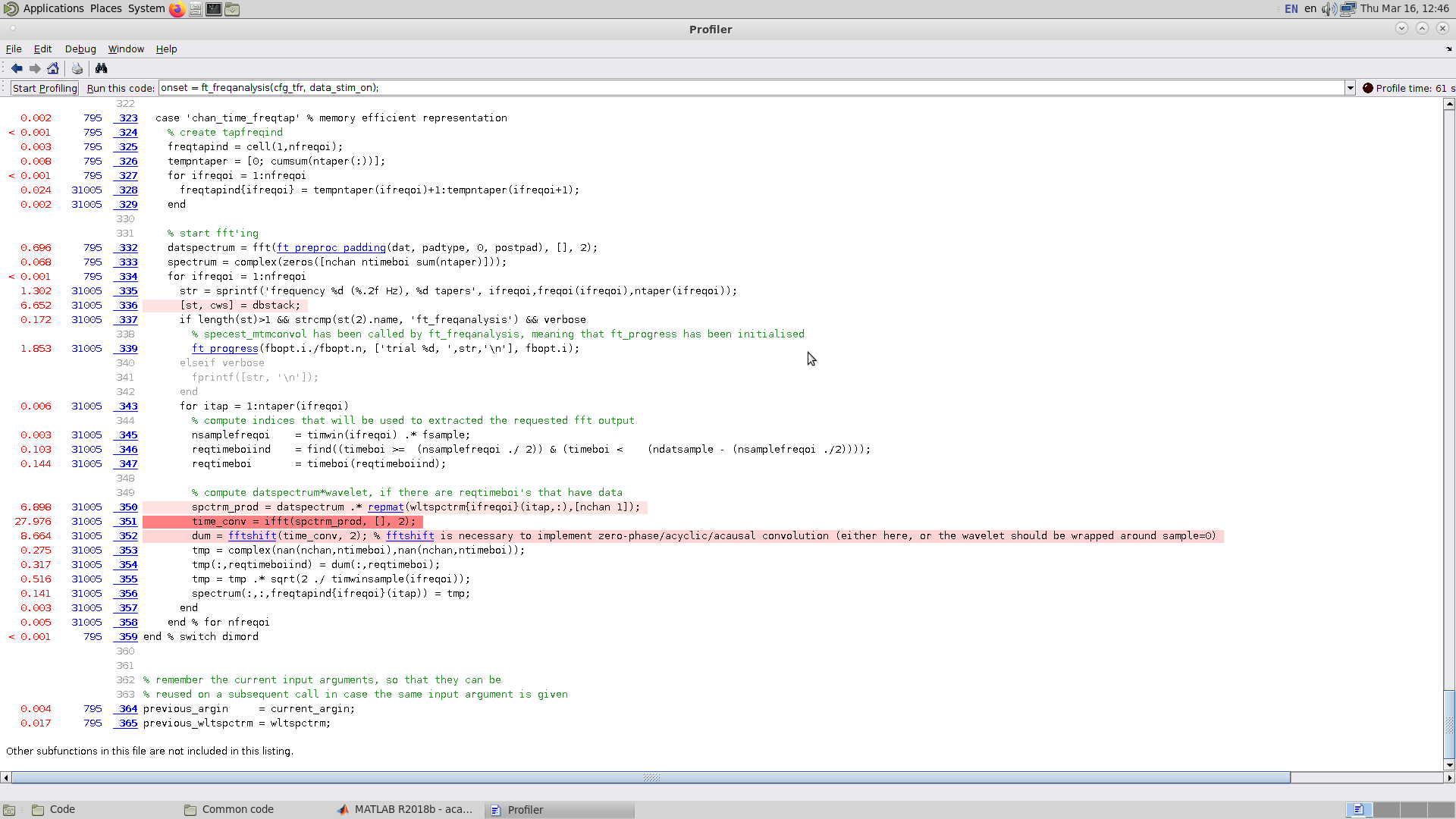
And the specific lines where most time is spent (I broke the convolution into 3 lines to see how long each step takes):
There are a few lines which take the vast majority of the run time:
Describe the solution you'd like
Here's what I think can be changed:
Adding a 'symmetric' flag to ifft() calls can speed it up, since the input data should always be real.That won't work because the data is real but the wavelets are complex.Do my suggestions make sense? I'd be happy to work on a PR.
The text was updated successfully, but these errors were encountered: